
Learn to train RF-DETR for precise segmentation of bacterial colonies on agar plates with edge deployment on NVIDIA Jetson.
Traditional auto-annotation tools like V7's Auto-Annotate rely on edge detection and color contrast. While effective for everyday photography, they struggle catastrophically with medical imaging, where subtle pathologies often have minimal contrast against surrounding tissue. A gray tumor on gray lung tissue? Edge detection fails.
Roboflow’s RF-DETR takes a different approach: transformer attention mechanisms that learn spatial patterns rather than hunt for edges. The result is accurate segmentation even when traditional computer vision can't find boundaries.
Traditional segmentation models struggle with a trade-off: accuracy or speed. Heavy transformers nail the complex, irregular boundaries of tumors and bacterial colonies, but they're painfully slow. Lightweight models are fast but miss critical details.
RF-DETR breaks this compromise. It brings transformer-level understanding to medical imaging while maintaining real-time inference speeds.
The result? You get pixel-perfect masks that trace the jagged edges of bacterial colonies, the kind of precision required for clinical accuracy, while still being fast enough to deploy on edge devices like NVIDIA Jetson.
This is what makes it perfect for medical imaging: the accuracy you need with the speed required for production deployment.
Medical Imaging Workflow with Roboflow: From Petri Dish to Production
In medical imaging, precision isn't just a nice-to-have, it's critical. A misidentified bacterial colony could mean the wrong antibiotic prescription. A missed tumor on a pathology slide could delay life-saving treatment. Research shows that each four-week delay in cancer treatment increases mortality risk by 6-8% for surgical patients, making speed and accuracy in diagnosis critical. Traditional computer vision approaches struggle here because medical images present unique challenges: specialized imaging equipment, domain-specific context, and the need for pixel-perfect accuracy on subtle features.
This is where RF-DETR shines. Unlike foundation models trained on everyday photos, RF-DETR can learn the nuanced patterns in medical imagery, whether that's segmenting bacterial colonies on agar plates or tracing irregular tumor boundaries in tissue samples. Let's walk through the complete workflow using a real clinical microbiology use case.
Step 1: Getting Your Data Into Roboflow
Before we dive into the bacteria colony dataset, here's the quick setup process if you're bringing your own medical images.
- Navigate to your Roboflow workspace and click Create New Project.
- You'll fill in a few key fields:
- project name (e.g., "Clinical Microbiology Detection")
- annotation group (your class names like "E-coli" or "S-aureus")
- visibility settings
- license information
- project type (select Instance Segmentation for pixel-level masks)
Once created, you'll land on the upload interface where you can drag and drop your images. Roboflow supports various image formats, including JPG, PNG, and others, making it straightforward to work with microscopy images or converted medical scans.
For this tutorial, we'll use the pre-existing Bacteria Colonies Detection dataset from Roboflow Universe. This dataset contains ~10,000 high-quality images of bacterial cultures on agar plates, a perfect representation of real clinical microbiology workflows.
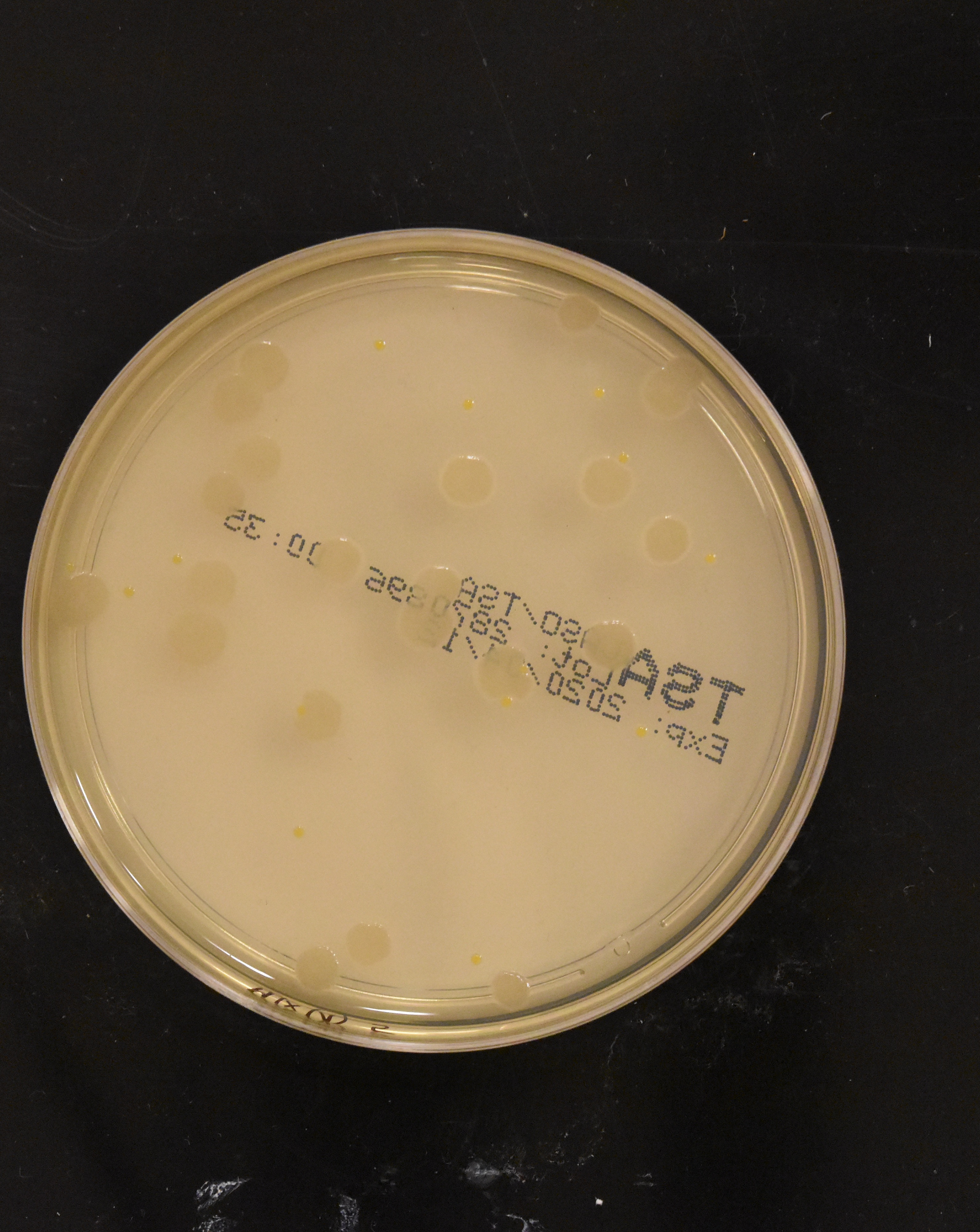

Bacterial colony detection is a genuine clinical need. Hospital labs process numerous culture plates daily, manually counting colonies to diagnose infections and test antibiotic resistance. The process is tedious and error-prone - technicians squint at plates under bright lights, marking colonies with markers to avoid double-counting. Each plate requires several minutes of focused counting. Automating this with computer vision means faster diagnoses and reduced human error, exactly the kind of high-impact medical application edge AI enables.
Step 2: SAM 3 Auto-Annotation
This is where the magic happens. Roboflow's SAM 3 Auto Label feature can intelligently segment objects across your entire dataset with just a text prompt.
If you're using the tutorial dataset: Since we forked an object detection dataset with bounding boxes, we need to start fresh with segmentation masks. Navigate to the Annotate tab and move the dataset to "Unassigned" to clear the existing labels. Then click "Annotate Images" followed by "Auto-Label Entire Batch."
If you uploaded your own images: After upload, click "Save and Continue," then immediately select "Auto-Label Entire Batch" to begin the auto-labeling process.
Now comes the crucial part: guiding SAM 3 with visual descriptions. For this workflow, we'll create a single "bacteria" class rather than distinguishing individual species. This mirrors real clinical practice - technicians first identify colonies present, then perform species-specific tests (like gram staining or biochemical assays) on promising candidates. The model handles the high-volume initial detection, letting lab staff focus their expertise where it matters most.
In the visual description field, enter:
white circle, brown spot, orange colony, round growth, circular colonyThis prompt tells SAM 3 to look for circular objects in various colors, capturing the diversity of bacterial species (Bacillus appears cream-colored, E. coli can show pink, Pseudomonas has green-blue pigment, Staphylococcus displays golden yellow). By specifying shapes (circle, round, circular) and colors (white, brown, orange), we're teaching SAM 3 to recognize the visual pattern that unites all colonies, distinct circular growth, while accommodating the color variations that distinguish different species.

Click Generate Test Results, and SAM 3 will process four sample images, giving you a preview of detection quality. Adjust the confidence threshold until the model accurately captures bacterial colonies without detecting the petri dish borders or background artifacts. In this case, a threshold around 40-50% worked well.

Once satisfied, auto-label your batch. SAM 3 processes up to 500 images at a time, segmenting each batch in minutes, a task that would take days of manual annotation. For the full 10k dataset, you'll repeat this process across multiple batches. Review the results in the Annotate view and make any necessary corrections.
Step 3: Roboflow's RF-DETR in Action
With the dataset labeled, we're ready to train. Navigate to the Versions tab and click Create New Version.
Configure your dataset split: 70% train, 20% validation, 10% test. This split is optimal for datasets in the 500-image range, providing enough training data while maintaining robust validation and test sets for reliable metrics.

Once your version generates, hit Train Model and select RF-DETR Small.
Why Small? With a single class and 500 images, Small provides excellent accuracy while maintaining fast inference speeds, critical for edge deployment. Larger models would offer marginal gains at the cost of deployment efficiency.
Training wraps up in about 30 minutes. The results on bacterial colony segmentation:
- mAP@50: 93.3% - The model achieves precise masks on most colonies
- Precision: 90.3% - Few false positives (detecting non-colonies)
- Recall: 87.1% - Captures the vast majority of bacterial growth

These metrics represent production-ready performance for clinical lab automation.
Step 4: From Cloud to Clinic
Here's where RF-DETR's edge deployment capabilities become game-changing for medical applications. While cloud-based inference works for research, clinical settings demand different constraints: patient data privacy (HIPAA compliance), low-latency processing for time-sensitive diagnoses, and offline capability in areas with unreliable connectivity.
Traditional platforms like V7 rely heavily on cloud infrastructure, requiring patient data to travel off-premises for inference. This creates compliance headaches, introduces latency, and makes offline operation impossible.
NVIDIA Jetson Orin is purpose-built for medical edge AI. Deploy your trained RF-DETR model directly to Jetson hardware stationed in hospital labs. Technicians can photograph culture plates, get instant colony counts and species identification, and receive results without patient data ever leaving the facility.
The workflow becomes: culture plate → smartphone photo → local Jetson inference → immediate diagnostic feedback. In practice, this means a lab technician photographs a completed culture plate using a mounted smartphone or camera. The image feeds directly to the Jetson device running your RF-DETR model. Within seconds, the system overlays segmentation masks on each colony, provides a total count, and can flag unusual colony patterns for human review. No cloud round-trips. No data exposure risks. Just fast, private, accurate medical AI where it's needed most.
This is the future of clinical automation, and RF-DETR makes it accessible without requiring massive labeled datasets or cloud infrastructure dependencies.
Conclusion
You've just built a complete medical imaging pipeline, from raw petri dish photos to a production-ready segmentation model. More importantly, you've done it in hours, not weeks, and without needing thousands of hand-labeled images or a team of ML engineers.
This workflow represents a fundamental shift in medical computer vision. Traditional approaches required either massive labeled datasets or cloud-based foundation models that couldn't handle domain-specific medical patterns. RF-DETR bridges that gap. It learns the subtle visual features that matter in clinical settings, whether that's distinguishing colony morphology, identifying tissue abnormalities, or tracing anatomical structures, and it does so with datasets you can actually collect and label.
The edge deployment capability seals the deal. Clinical environments need models that process patient data locally, maintain low latency for time-sensitive diagnoses, and operate reliably without constant internet connectivity. By deploying RF-DETR on NVIDIA Jetson hardware, you're delivering all three.
The bacterial colony detector you've built isn't just a proof of concept. It's a template for real clinical automation: culture plate imaging systems that provide instant colony counts, reduce technician workload, and accelerate antibiotic resistance testing. That's the promise of edge AI in healthcare, and with RF-DETR and Roboflow, it's finally accessible.
Further reading
Below are a few related topics you might be interested in: